Most people assume the human genome is a fully decoded library. One complete blueprint, every page accounted for. The reality is stranger and more fascinating. Decades after the first draft of the human genome was published, substantial portions of our DNA remain poorly understood, technically difficult to read, and in some cases, not even recognizably human in origin.
These are the genetic ghosts. Ancient viral passengers, repetitive regions that confuse our best machines, and stretches of sequence so structurally complex that they resisted mapping until just recently. Understanding them is becoming one of the most urgent frontiers in genomics.
The Missing Percent: What the Human Genome Project Left Out

The Human Genome Project, launched in 1990 and declared complete in 2003, was a landmark achievement. It was declared complete on 14 April 2003, and included about 92% of the genome. That remaining fraction wasn’t a minor footnote. That historic draft, and subsequent human DNA sequences, all missed about 8% of the genome.
The standard sequencing technology works fine for most of the genome, but not in areas where DNA code is the result of long repeating patterns. If a supercomputer only had small fragments, how could it assemble a DNA sequence that repeated “AGAGAGA” for bases upon bases? That’s what the missing 8% of the genome looked like.
Large stretches, totaling nearly 200 million base pairs, roughly the size of all of chromosome 3, remained unresolved and were represented by gaps or model sequences. This wasn’t a small rounding error. It was a continent-sized blank on the map of who we are.
Dark Regions: DNA That Standard Technology Cannot Read
The human genome contains “dark” gene regions that cannot be adequately assembled or aligned using standard short-read sequencing technologies, preventing researchers from identifying mutations within these gene regions that may be relevant to human disease. Researchers have since classified these into two main types. There are regions with few mappable reads, called dark by depth, and others that have ambiguous alignment, called camouflaged.
Based on standard whole-genome Illumina sequencing data, researchers identified 36,794 dark regions in 6,054 gene bodies from pathways important to human health, development, and reproduction. Of these gene bodies, roughly nine percent are completely dark and more than a third are at least five percent dark.
A large number of gene bodies, some 527, were one hundred percent dark, which means they are entirely overlooked in standard whole-exome, whole-genome, and RNA sequencing studies. Think about that for a moment. Entire genes, invisible to the tools most labs use every day.
The Dark Proteome: Over One-Third of Protein-Coding Genes Still Unknown
The biological functions and disease relevance of the “dark genome,” which includes over one-third of all protein-coding genes, remain largely unknown. Researchers are now using integrative network and functional analyses to construct a systems-level map of dark gene contributions to human genetic diseases.
Scientists have identified 16 hub dark genes, including R3HDM2 and RPUSD4, that are central to disease networks and are overwhelmingly enriched for roles in mitochondrial protein synthesis. These hubs form modular networks connecting major inflammatory conditions like psoriasis and tuberculosis.
Furthermore, the expression of these hubs is controlled in a tissue-specific manner by thousands of genetic variants, providing direct mechanistic links to phenotypes such as myocardial infarction and diabetes. In other words, the dark genome isn’t just mysterious. It’s clinically relevant in ways we’re only beginning to measure.
Ancient Viruses Inside Us: The Story of Human Endogenous Retroviruses
The retroviruses that infected our ancestors inserted copies of viral DNA into the genome, and these sequences were copied to allow for the viruses to replicate. This viral DNA has been passed down through generations, and now the human genome is littered with hundreds of thousands of integrations of endogenous retroviruses, amassed over millions of years. They are part of the so-called “dark genome,” sections of DNA that also include other diverse and often larger families of transposable elements.
Human endogenous retroviruses comprise approximately 8% of the human genome, serving as fragments of ancient retroviral infections. To put that in perspective, approximately 8% of the human genome, over four times more than its protein-coding regions, comprises sequences of viral origin.
Genomic analyses have revealed that HERV sequences comprise 8% of the human genome, with approximately 98,000 annotated insertions. Most of these are degraded remnants. Most of these have acquired inactivating mutations during host DNA replication and are no longer capable of producing the virus. Still, some remain active in ways that surprise researchers.
Not Just Junk: What These Viral Fossils Actually Do

It was long thought that HERVs were “junk DNA.” However, it is now known that HERVs are involved in various biological processes through encoding proteins, acting as promoters and enhancers, or long non-coding RNAs that affect human health and disease.
While these DNA sequences are parasitic in origin, they are now responsible for at least part of what makes us human. Research from the Crick Institute highlights striking examples of this integration. Recombinases derived from retroviral elements function as naturally evolved gene editing tools, giving the immune system its remarkable adaptability. Even the evolution of the placenta and the loss of the tail in human ancestors have connections to these ancient viral genes.
HERVs account for between 0.1 and 0.4 percent of all translation in distinct tissue-specific profiles. Collectively, these findings support claims that HERVs are actively translated throughout healthy tissues, contributing sequences of retroviral origin to the human proteome. These are not silent fossils. They remain at work inside every human body.
When the Ghosts Wake Up: HERVs and Disease
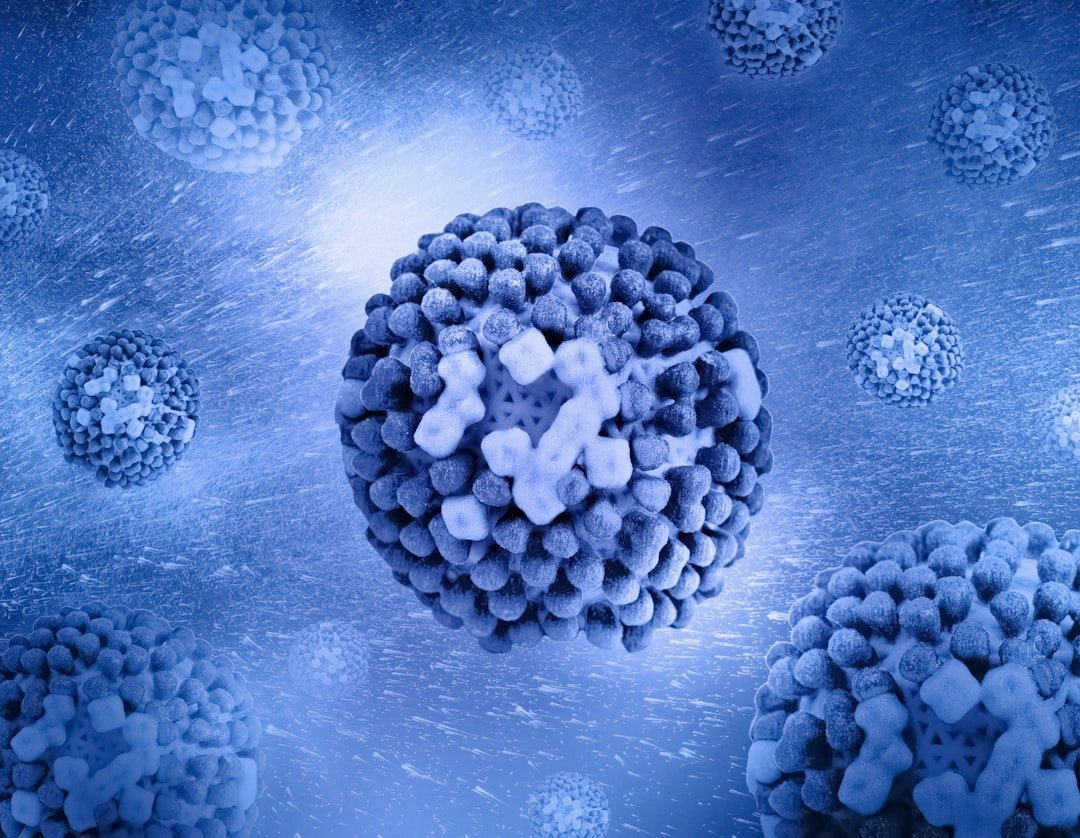
Human endogenous retroviruses are generally maintained in a silenced state by robust epigenetic mechanisms. However, specific HERV groups, particularly HERV-W and HERV-K, can become derepressed under specific pathological conditions, thereby contributing to the initiation and progression of neuroinflammatory and neurodegenerative processes.
In some cells, including cancer cells, researchers have observed immune responses to these endogenous retroviruses. This is because, in cancer, the body effectively loses control of much of the genome, including these repetitive viral elements. This means that formerly repressed DNA sequences are transcribed, sparking excitement about the potential of the newly expressed proteins as targets for cancer immunotherapies.
The HERV-K envelope glycoprotein is aberrantly expressed in cancers, autoimmune disorders, and neurodegenerative diseases, and is targeted by patients’ own antibodies. A 2025 paper published in MDPI Vaccines noted that researchers are now investigating monoclonal antibodies designed to directly target these elements, representing a genuinely novel therapeutic strategy rooted in the dark genome.
Centromeres and Telomeres: The Last Unmappable Fortresses
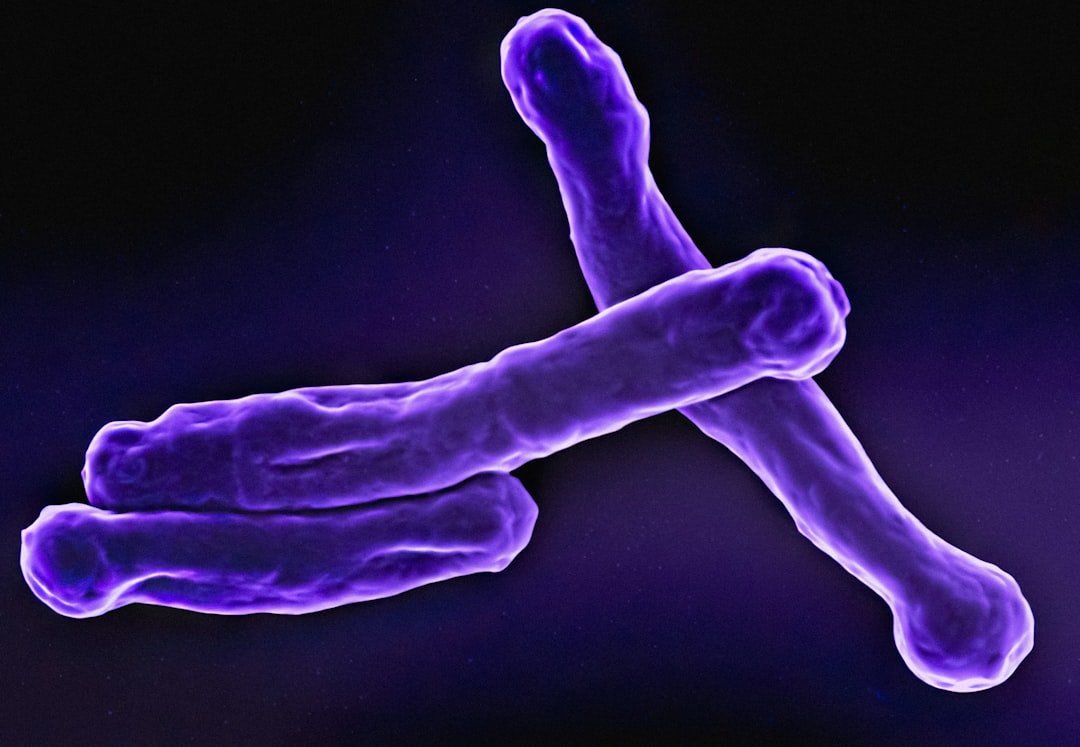
Among the “unmappable” regions were some of the most recognizable structures in biology. Those knots visible on chromosomes are centromeres, bundles of DNA that hold the chromosomes together. They play a key role in cell division.
Centromeres are the chromosomal domains required to ensure faithful transmission of the genome during cell division. They have a central role in preventing aneuploidy by orchestrating the assembly of several components required for chromosome separation. However, centromeres also adopt a complex structure that makes them susceptible to being sites of chromosome rearrangements.
Telomeres, the protective caps at the ends of chromosomes, are crucial in maintaining genomic stability and cellular longevity. These specialized DNA-protein structures prevent chromosome degradation, fusion, and recognition as damaged DNA, thereby safeguarding the integrity of the genetic material. Telomere shortening serves as a biological clock of cellular ageing. Both regions were among the last to yield to sequencing technology and remain subjects of intense study today.
The T2T Breakthrough: Finally Reading End to End

Two major advances have emerged to address these shortcomings: complete gap-free human genome sequences, such as the one developed by the Telomere-to-Telomere Consortium, and high-quality pangenomes, such as the one developed by the Human Pangenome Reference Consortium. Facilitated by advances in long-read DNA sequencing and genome assembly algorithms, complete human genome sequences resolve regions that have been historically difficult to sequence, including centromeres, telomeres, and segmental duplications.
The most recent reference assembly, T2T-CHM13, was generated using a combination of PacBio HiFi and Oxford Nanopore ultralong-read sequencing and represents the first complete genome, including the 8% of the genome that had remained hidden since the first human reference genome was published in 2000.
Yet the work isn’t finished. Short-read sequencing technologies are inherently limited in their ability to resolve highly repetitive, structurally complex, and low-mappability genomic regions. Long-read sequencing technologies, such as PacBio and Oxford Nanopore Technologies, offer improved resolution of these regions, yet they are not perfect. Completing the map required not just better machines, but an entirely different approach to reading DNA.
The Dark Genome and Human Disease: A Direct Connection
The inability of traditional DNA sequencing methods to analyze genomic “dark regions” limits our understanding of the full genetic architecture of disease. Dark regions are highly repetitive elements that are not resolved via traditional sequencing, many of which are lacking in the GRCh38 human reference genome. These regions include low-complexity microsatellites, transposable element-rich sequences, centromeric DNA, and rDNA arrays.
A 2025 study published in Alzheimer’s & Dementia applied long-read sequencing to dark genomic regions in brain tissue from Alzheimer’s patients, revealing previously invisible genetic and epigenetic changes. These findings built upon long-read sequencing of DNA isolated from neuronal nuclei of control and late-stage Alzheimer’s disease brains, with researchers leveraging the telomere-to-telomere reference genome and methylation data to comprehensively interrogate genomic dark regions.
Advances in 3D genomics and multiomics are enabling scientists to uncover disease mechanisms hidden in the human genome’s noncoding regions. Around 98.5% of human DNA is non-coding, meaning it doesn’t get copied to make proteins. The vast majority of disease-associated mutations found in genome-wide studies fall precisely in these overlooked zones.
What Comes Next: Reading the Ghosts
Long-read sequencing technologies such as PacBio and ONT have been shown to reduce the amount of dark gene-body regions by up to 77%. That’s a meaningful reduction, though it still leaves a significant portion unresolved. The field is now moving toward pangenome references that capture diversity across populations, not just one reference individual.
Scientists have pieced together a new draft of the human genome that better captures humanity’s genetic diversity. The new “pangenome” incorporates the DNA of 47 individuals from every continent except Antarctica and Oceania. The new reference adds 119 million base pairs to the library of previously known base pairs, deepening our understanding of human genetic diversity and making it more complete.
An international conference on the hidden cell and the dark genome, organized in partnership with the Wellcome Discovery Research Platform and the MRC Human Genetics Unit at the University of Edinburgh, aims to provide a forum to present cutting-edge research, and to discuss how novel approaches and methodologies will make substantial breakthroughs in these critical areas of biology. The field is moving fast, and the ghosts in our genome are beginning to speak.
Conclusion

The idea that we have fully decoded the human genome has always been a simplification. The truth is that our DNA harbors entire evolutionary histories: viral remnants, ancestral debris, structurally complex repeats, and functionally vital regions that only recently became readable. These are not errors in the genome. They are a record of survival across deep time.
What researchers are now uncovering is that the dark genome is not peripheral to human biology. It sits at the center of immunity, development, aging, and disease. The more we learn to read these regions, the more it becomes clear that the least understood parts of our DNA may hold some of the most consequential answers. The ghosts were always there. We just didn’t have the right tools to see them.